In an experimental study from researchers across India, Singapore, and the USA, IQM’s Sirius 24-qubit superconducting processor was utilized to demonstrate high-accuracy molecular simulations. The research employed Sample-based Quantum Diagonalization (SQD), a noise-resilient paradigm that uses the quantum processor as a “configuration sampler” rather than a direct energy estimator. By identifying high-weight Slater determinants (electron configurations) on the hardware and performing exact diagonalization on a classical computer, the team decoupled the final energy precision from individual quantum gate errors, achieving results within chemical accuracy for several benchmark molecules.
The study benchmarked two distinct quantum circuit “blueprints” or ansätze: the Local Unitary Cluster Jastrow (LUCJ) and the Linear-CNOT Unitary Coupled-Cluster (LCNot-UCCSD). The LUCJ ansatz proved highly effective due to its shallow circuit depth, while the LCNot-UCCSD, despite reducing classical pre-computation needs, faced challenges with deeper circuits that impacted data recovery for larger molecules like water (H2O) and ammonia (NH3). This comparative analysis provides a roadmap for selecting the most efficient algorithms for “noisy” near-term quantum hardware.
A significant highlight of the paper is the generation of Potential Energy Surfaces (PES). The researchers reported the first experimental generation of a full two-dimensional (2D) PES map for the water molecule on superconducting hardware. By scanning a 32×32 grid of bond lengths and bond angles, they mapped the molecule’s energy landscape with precision matching exact classical full configuration interaction (FCI) results. These surfaces are critical for understanding how molecules vibrate, rotate, and undergo chemical reactions.
To address larger, more complex systems, the team integrated SQD with Density Matrix Embedding Theory (DMET). This hybrid strategy allows a large molecule to be broken down into smaller “fragments” that can be computed on the quantum hardware, while the rest of the molecular environment is handled classically. This allowed the researchers to simulate eight ligand-like molecules and the antiviral drug Amantadine, representing a significant step toward using quantum computers for pharmacological research and drug discovery.
The results establish a strong baseline for quantum-centric supercomputing, where quantum processors act as specialized accelerators within classical high-performance computing (HPC) workflows. By demonstrating that 16 operational qubits can provide chemically accurate data for real-world molecules, the study validates the reliability of current superconducting architectures and points toward future workflows capable of tackling classically intractable problems in materials science and catalysis.
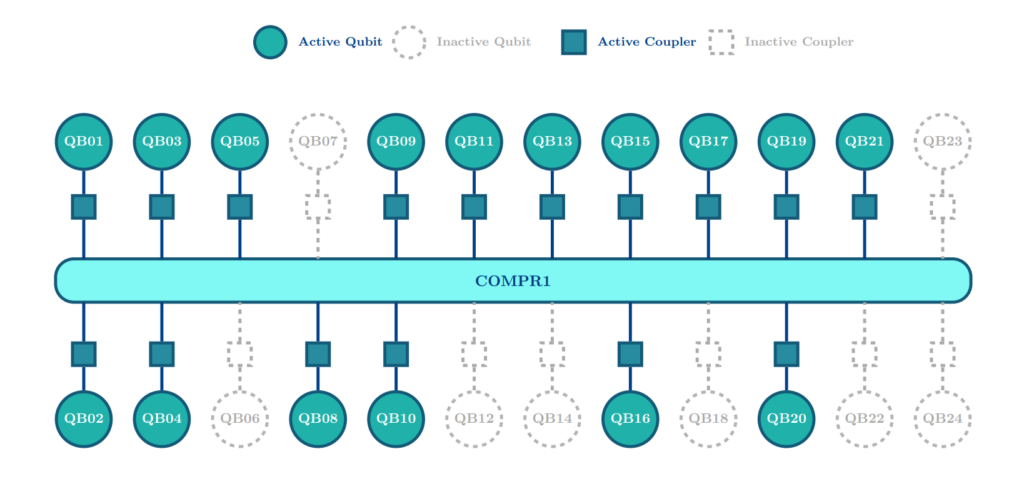
For the full technical details of this experimental demonstration on IQM hardware, access the paper on arXiv here. Information regarding the IQM Sirius processor and its star topology can be found here.
April 18, 2026